Originally posted at nunocruz.pt
Genome, like any other Feedzai product, is subject to an exhaustive battery of end-to-end (E2E) tests to ensure it works according to its design specifications. Since Genome is a highly visual tool, testing requires a very particular approach. We build the graph area using an HTML canvas element, which prevents us from using conventional E2E techniques. In this blog post, we'll explain how we approached and solved the problem of testing an application whose main interactions occur on a canvas element.
Testing a whiteboard
Testing a traditional web application (by "traditional" I mean a User Interface (UI) where each piece of information is displayed using HTML elements) is more or less straightforward with the current E2E frameworks like Selenium or Cypress. Overall, the workflow consists of performing a set of actions on the page, such as clicking on buttons or typing text on input elements and asserting the page gets into the desired state (e.g., asserting that some elements are present on the page).
This works well for almost every HTML element except the canvas element. If you are not familiar with canvas, it is an HTML element that can be used to draw graphics via scripting. We can imagine it as a whiteboard where you can freely draw anything.
Compared with other HTML elements where we can nest several elements inside each other (DOM tree) to produce content, what we see on a canvas element is not represented in any nested element, which means that it doesn't produce a queryable DOM tree. From a testing perspective, how can we assert that our application gets into the desired state if we cannot query the content of a canvas element? For example, how can we test that after deleting a node from a graph, we get n - 1 nodes?
Exposing an API
Before explaining how we implemented the E2E tests for Genome, it is important to give a little bit of context on how Genome, and particularly its graph, is built.
Genome's graph is built using a combination of React and D3.js. A React component named GenomeGraph embeds the canvas element. Overall, this component is responsible for setting up and handling user events that target the graph. It also communicates with D3.js to compute the nodes and edges positions.
D3.js is a well-known library that is used to help build visualization tools. With Genome, by default, we use the D3.js force-directed graph layout. This layout simulates physical forces on nodes until they balance (their final position). Visually speaking, the more simulations, the more the nodes tend to separate from each other, while fewer simulations might make them stay closer to each other. Figure 1 illustrates a scenario where the number of simulations is greater than the one in Figure 2.
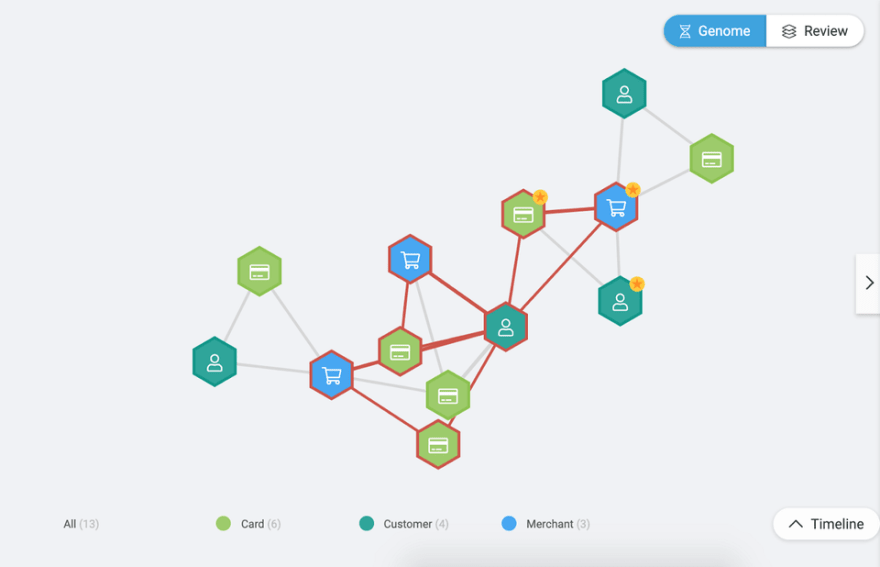 Figure 1. D3.js force simulation with around 300 iterations. As explained, in this case, the nodes are slightly apart from each other when compared with the graph in Figure 2.
Figure 1. D3.js force simulation with around 300 iterations. As explained, in this case, the nodes are slightly apart from each other when compared with the graph in Figure 2.
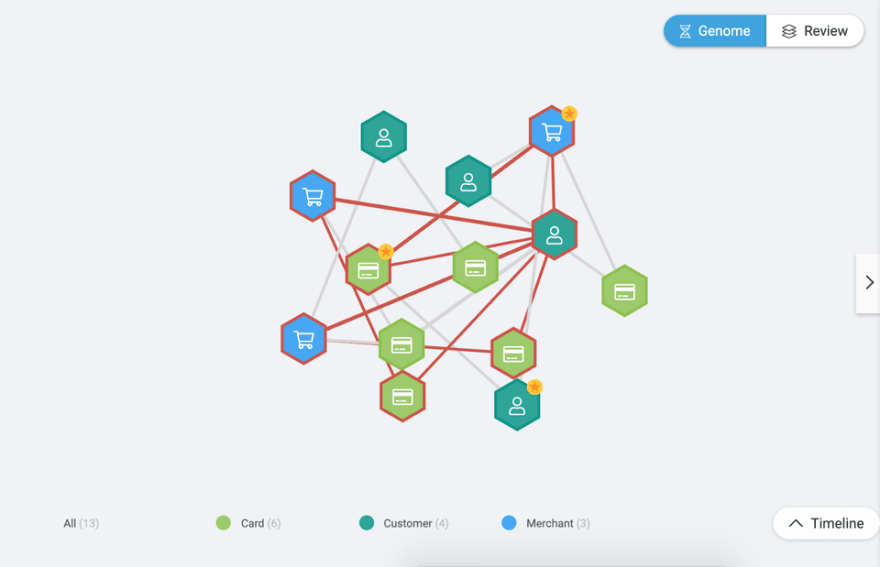 Figure 2. D3.js force simulation with around 13 iterations.
Figure 2. D3.js force simulation with around 13 iterations.
Cypress over Selenium
We decided to use Cypress to implement our E2E tests. Why Cypress and not Selenium? Cypress is more developer-centric, in the sense that any Frontend developer can easily implement E2E tests using JavaScript. Another advantage is that where Selenium WebDriver runs remotely outside the browser, Cypress is the exact opposite; it runs inside the browser. The ability to run code inside the browser simplifies things. For example, you can simply drop a debugger into your application or spec code - something that makes it super easy to use the developer tools while you're developing. Besides the technical aspect, when we made our decision, Cypress was getting a lot of traction in the Frontend community. Taking all of this into account, we decided to give Cypress a try (no regrets so far).
Now that we have more context on the tools used to implement and test the Genome graph, it's time to detail how we were able to test a graph drawn in a canvas element.
How to test a graph drawn in a canvas element
After some analysis, we decided to use an API to expose the Genome application for consumption by our Cypress application. This allowed us to manipulate the graph and assert its state. Due to the characteristics of the GenomeGraph component, it was a "natural" choice for us to use it to help build the API.
A High-Order Component (HOC) is an advanced technique in React for reusing component logic. We used this technique because it generates an enhanced component from a given component. In particular, we produced an enhanced component from the GenomeGraph component with the ability to create an API and expose it in the browser's global object. Even though it's not reusable, this approach seemed the cleanest and most elegant way to expose a component's API.
Since we didn't want this API exposed in the production environments, we defined a variable to enable it only in testing environments.
From the set of operations that the API exposes, the most useful one is the waitUntilGraphSimulationEnd operation. It allows us to wait for the D3.js simulation to finish in order to start interacting with the graph. Without it, it would be very difficult to interact with the graph and would probably lead to flaky tests because the graph was still "moving" from its initial to its final state.
The snippet below describes the implementation of the function that waits for the simulation to end.
Implementing custom commands
From the perspective of the Cypress application, we created several custom commands to abstract the communication with the Genome graph. The snippet below describes the implementation of the getGraphNodeInformationById and the waitUntilGraphSimulationEnd custom commands.
While implementing our tests, we use the custom commands as if they were part of the Cypress library. The example below is a test that asserts that when clicking on a specific node, that node gets selected. You can notice the usage of the previously mentioned waitUntilGraphSimulationEnd and getGraphNodeInformationById custom commands.
Final thoughts
Adopting an approach based on "direct communication" between the Cypress application and the Genome application allowed us to include our main component (Genome graph) in the set of E2E tests. However, we are aware of the drawbacks of this approach.
One of the goals of E2E testing is to interact with the application like a user would: click on the screen, input text and expect to see changes on the screen. By using an API-based approach, we somehow break this goal. In practice, we fake the interaction with the graph by calling code directly from the Genome application.
Another limitation are the drag and drop operations. Currently, we cannot test the scenario of dragging nodes to some part of the screen because we noticed that it introduces flakiness in our tests. While we are working on a solution to overcome this limitation, a set of manual tests were added to cover the drag and drop operations.
Visual testing, an alternative to functional testing, is an approach that should be explored in the near future. This technique takes image snapshots of the application and compares them with previously approved baseline images. It is a technique that fits well on features that mainly encode information through visualization. For example, in Genome, we have a new feature named Lenses which aims to apply different colors to nodes and edges to encode certain information, e.g., entities that participated in, at least one fraudulent transaction are displayed in red, while entities participating only in genuine transactions are displayed as green. Comparing image snapshots with baseline images would be an interesting way to assert that the lenses are correctly applied to the nodes and edges.
Our solution to test an application that is built mainly using canvas is far from perfect, and our goal is to continually iterate in order to solve the current limitations.
Note: The credit for this solution goes to Liliana Fernandes and Victor Fernandes. I'm only the messenger :D


Top comments (0)