In this article I will demonstrate how agent-based modeling can be used to model and investigate disease spread. I will use the abm_framework in Python to implement an agent-based SIR model in Python. This application was first published here: Agent-based SIR model (Python)
Introducing the SIR model
The SIR modeling framework is a popular framework for modeling disease spread. SIR stands for Susceptible, Infected, Recovered. Below figure illustrates the SIR model.
For example, anyone in the susceptible state can be infected. Once you are infected you can infect anyone in the susceptible state. The risk of infection can be modelled to be between 0% and 100%.
Anyone infected will recover with a certain probability. Once you are recovered, you are either resistant or you can fall back in to susceptible mode. Every state transition is described with probability ratios.
Introducing agent-based modeling in Python
Agent-based modeling is a popular simulation technique, next to discrete-event simulation, monte-carlo simulation and system dynamics modeling.
Agent-based models describe agents and their inner attributes and dynamics, as well as the interaction between agents. From this one hopes to study and understand the emergence of complex macroscopic system behaviour.
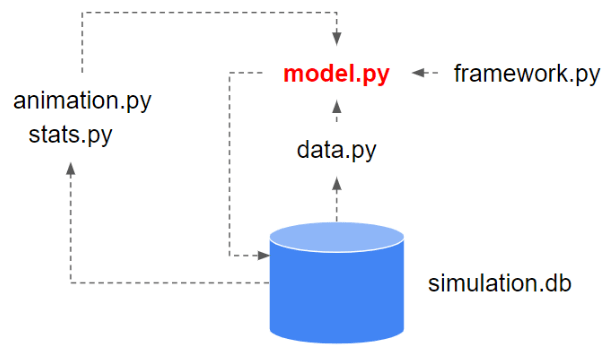
I use the abm_framework in Python for implementing agent-based simulation models. Its setup is illustrated in below figure.
model.py is the application specific model. It consumes the modules provided by the abm_framework. I implemented the SIR model in Python that way. You can find the code here. The model has the following specifications: 5% initial infection, 7% infection risk, 3% recovery chance. Agents interact with a defined neighbourhood, with a fix radius. Infected agents can infect other agents within their limits of that neighbourhood.
Simulation results
Below chart summarizes a simulation run with the parameter settings described in previous section.
Recovery and social distancing (limited infection radius) end disease spread. Only 12% of all agents are infected.









Top comments (0)