Reason 1: Esc twice ain’t it
I’m keyboard-driven, I use a tiling window manager, my mouse is seen as a luxury and I find it a pain to lift my hands from my keyboard and break my stream of thought to shift what feels like a 16 hour flight with 2 connections all the way to my mouse. (And it isn’t that I have a bad mouse, I like my mouse! But I like my keyboard more...).
For those who don’t know what a tiling window manager is, it’s essentially a way of interacting with your open windows, grouped into workspaces (which are switched by key-combinations). I like the idea of auto-aligning open windows (trust me, I’ve been through Alt-Tab hell and back), and having my applications open by keystrokes and automatically fill exactly a given part of the screen is a Godsend!
Everything you're seeing happens with TWO keystrokes
However, my biggest gripe with the entire notebook environment isn’t necessarily related directly to notebooks: it has to do with actual, WORKING vim-shortcut support.
I love shortcuts, and my entire being is wired to hit j and k as soon as I see any sort of code-related text to navigate up and down. My caps-lock key is remapped to Esc so that I have more control switching back to ‘Normal’ mode in vim. Usually this wouldn’t be an issue, since I use the vim-extension in VSCode and use evil-mode in emacs (I use doom-emacs which use vim-keybindings by default).
However, God forbid that you happen to hit the esc key twice whilst in a Jupyter/IPython cell with the Vim extension loaded (either via the web interface or via VScode), and you're taken OUT of the entire editor. This means that I now have to reach (what feels like) halfway across the room to my mouse, to re-click the cell I was working on, this is just terrible for ergonomics.
Now, this may not be an IPython-specific issue, it's just that I haven't found an extension which works, since technically speaking all the extensions work as they are supposed to! Hitting esc takes you to "normal" mode in Vim, however this now breaks the ability to shift between cells normally as you would in a Notebook because you technically aren't in the actual cell
Reason 2: print(type(df)) is NOT debugging
An IPython/Jupyter notebook is meant to be run sequentially, which I may or may not have an issue with. My MAIN gripe around notebooks however come in the form of debugging. Typically, a break-point is set at a particular line in code, and a debugger temporarily halts code execution at that point, and we can set certain variables to be "watched"; i.e. the debugger can keep track of these variables during execution. For example, in the following code
output = tf.nn.Conv2D(some_image, filter, ...
I'm applying a convolution to what is presumably some sort of image. Don't worry about the actual operation, the important part is that I'm taking an image (or a matrix of floating-point values) and applying some operation to it, which may change the actual values, change the type of values (some may go to NaN) and possibly change the actual shape of the original image depending on convolutions. Now, in order to actually check what the operation is doing, I could potentially do the following set of abominations:
print(output.shape) # get shape
print(type(output)) # make sure I actually get a return Tensor
print(np.isna(np.sum(output.values))) # make sure no NaNs popped up
and probably the WORST of them all:
# I'm PRINTING an IMAGE as its
# RAW values, how in the
# actual heck is this helpful..?
print(output)
Also to the above, remember IPython notebooks length grow as cell-outputs are populated. Imagine doing the print(output) but MULTIPLE times in one notebook just to validate pre-processing steps (which is very common in computer vision). This is just highly unproductive!
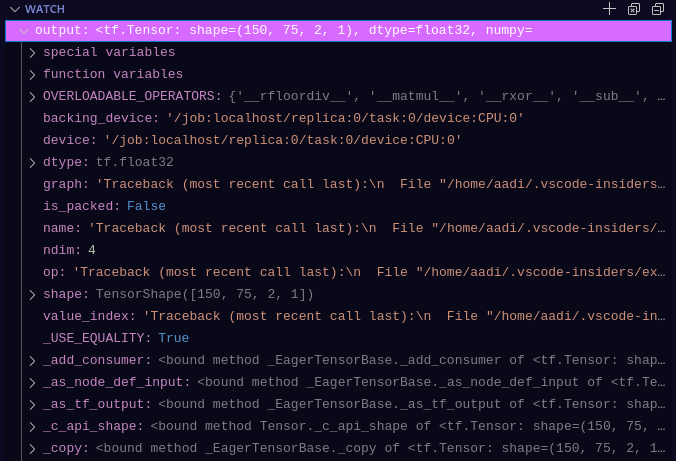
Compare this to the below which requires none of the print statements (which usually have to be removed or re-added every-time you need to find the values or properties of an object, which is tiring, unnecessary and overall unproductive). EVERYTHING I could ever want to know about any variable is seen, and is much more useful and clean-coding practice:
This is from using watch in VSCode's debugger on the output variable, the shape, type, etc etc are all visible without additional code
Finally: Becoming familiar with how an Engineer might look at your code to deploy
This may be the least "ranty" reason. Deployable code does NOT contain random print statement throughout, and typically forms a directed acyclic graph between functions! This acyclicity is broken in a Jupyter notebook, since changing a cell above another cell does NOT change the output of the below cell. Let me show you what I mean, if I do the following:
The output is quite clearly correct,however if I NOW change the value of a (or b) and re-run only the upper cell, clearly the following does not remain true:
Hence the entire notebook need-be re-run (which is not a big deal, there's a convenient drop-down, but then I have to use my mouse again AND it breaks the natural flow of functions throughout my pipeline). But the biggest issue I have here is modularity, I can't easily swap in one notebook for another without doing ALL the data pre-processing steps in separate cells. You can't import one notebook into another the same way you'd import packages.
And deployment?! YIKES, that's a data engineer's job right....? WRONG!! How about we actually think about how our code is to be deployed, and follow some sort of paradigm where our resulting code can be modularly swapped in and out, and most importantly, actually follow some format to be easily tested. There are frameworks that assist in setting up your entire pipeline as a directed-acyclic-graph (such as Kedro, etc), but it's still a chore, and highly inefficient to not consider that building a data-science pipeline, outside of purely academic research is not for deployment in some setting
In conclusion, there are benefits to IPython notebooks, ease of demonstration, etc etc. However, this article covers my opinion of Jupyter/IPython notebooks, and why I try my best to steer clear of them for data-science/machine-learning related tasks.
If you like rants like these, feel free to follow me on twitter









Top comments (0)